- HOME
- Enzyme List
- PNP-311 PURINE-NUCLEOSIDE PHOSPHORYLASE
PNP-311
PURINE-NUCLEOSIDE PHOSPHORYLASE from Microorganism
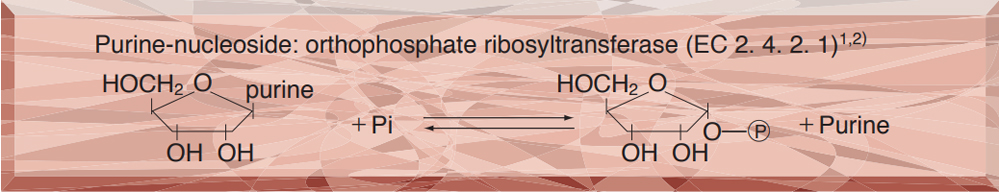
PREPARATION and SPECIFICATION
| Appearance | White amorphos powder, lyophilizedWhite amorphos powder, lyophilized | |
|---|---|---|
| Activity | GradeⅢ 15 U/mg-solid or more | |
| Contaminants | Catalase | ≤20 % |
| 5'-Nucleosidase | ≤1.0x10-3 % | |
| Adenosine deaminase | ≤1.0x10-3 % | |
| ATPase | ≤1.0x10-2 % | |
| Stabilizers | K-Gluconate, mannitol, EDTA | |
PROPERTIES
| Stability | Stable at −20 ℃ for at least one year (Fig.1) |
|---|---|
| Molecular weight | approx. 120,000 |
| Isoelectric point | 4.1±0.1 |
| Michaelis constant | 6.4×10-5 M (Inosine), 3.2×10-4 M (Pi) |
| Inhibitors | p-Chloromercuribenzoate, SDS, Hg2+, Ag+ |
| Optimum pH | 7.5−8.0 (Fig.3) |
| Optimum temperature | 65 ℃ (Fig.4) |
| pH Stability | pH 6.0−9.0 (30 ℃, 16 hr) (Fig.5) |
| Thermal stability | below 60 ℃ (pH 7.7, 30 min) (Fig.6) |
| Substrate specificity | (Table 1) |
| Effect of various chemicals | (Table 2) |
APPLICATIONS 3~5)
This enzyme is useful for enzymatic determination of inorganic phosphorus, 5'-nucleotidase and adenosine deaminase, in combination with xanthine oxidase (XTO-212) and uricase (UAO-201, UAO211)
ASSAY
Principle

The formation of uric acid is measured at 293 nm by spectrophotometry.
Unit definition
One unit causes the formation of one micromole of uric acid per minute under the conditions detailed below.
Method
Reagents
| A. K-Phosphate buffer, pH 7.7 | 50 mM |
|---|---|
| B. Inosine solution | 32 mM[Dissolve 85.8 mg of inosine (MW=268.23) in 10 mL of H2O with heating] (Stable for at least two weeks if stored at 4 ℃) |
| C. Xanthine oxidase solution | ca.6.6 U/mL[Dissolve xanthine oxidase (XTO-212) to ca.6.6 U/mL with ice-cold buffer A](Should be prepared fresh) |
| D. Enzyme diluent | buffer A |
Procedure
1.Prepare the following reaction mixture in a cuvette (d = 1.0cm) and equilibrate at 37 ℃ for approximately 5 minutes
| 2.7 mL | Potassium phosphate buffer, pH 7.7 | (A) |
|---|---|---|
| 0.2 mL | Substrate solution | (B) |
| 0.1 mL | Xanthine oxidase solution | (C) |
| Concentration in assay mixture | |
|---|---|
| Potassium phosphate | approx. 47 mM |
| Inosine | 2.1 mM |
| Xanthine oxidase | approx. 0.2 U/m |
2.Add 0.05 mL of the enzyme solution* and mix by gentle inversion.
3.Record the decrease in optical density at 293 nm against water for 3 to 4 minutes with a spectrophotometer thermostated at 37 ℃, and calculate the ΔOD per minute from the initial linear portion of the curve (ΔOD test).
At the same time, measure the blank rate (ΔOD blank) using the same method as in the test except that the enzyme diluent is added instead of the enzyme solution.
*Dissolve the enzyme preparation in ice-cold enzyme diluent (D), and dilute to 0.1−1.5 U/mL with the same buffer and store on ice.
Calculation
Activity can be calculated by using the following formula :
Volume activity (U/mL) =
-
ΔOD/min (ΔOD test−ΔOD blank)×Vt×df
12.5×1.0×Vs
= ΔOD/min×4.88×df
Weight activity (U/mg) = (U/mL)×1/C
| Vt | : Total volume (3.05 mL) |
| Vs | : Sample volume (0.05 mL) |
| 12.5 | : Millimolar extinction coefficient of uric acid under the assay condition (cm2/micromole) |
| 1.0 | : Light path length (cm) |
| df | : Dilution factor |
| C | : Enzyme concentration in dissolution (c mg/mL) |
REFERENCES
1) R.E.Parks, Jr. and R.P.Agarwal; The Enzymes,Vol.7, p483 (3rd ed.)(1972)
2) P.A.Hoffe, R.May and B.C.Robertson; Methods in Enzymology,Vol.11, p70 (1972)
3) Y.Machida and T. Nakanishi; Agric.Biol.Chem.,45, 1801 (1981)
4) M.Sugiura, K.Kato, T.Adachi, Y.Ito, K.Hirano and S.Sawaki; Chem.Pham.Bull.,29, 1451 (1981)
5) P.Fossati; Analytical Biochemistry.,149, 62 (1985)
Table 1. Substrate Specificity of Purine-nucleoside phosphorylase3)
[Inosine: Purine-nucleoside phosphorylase - Xanthine oxidase system, pH 7.7 Guanosine, Adenosine, ATP, Thymidine: UV-system, pH 7.4]
-
Substrate(0.2mM) Relative activity(%) Inosine 100 Guanosine 41 Adenosine 0 -
Substrate(0.2mM) Relative activity(%) ATP 0 Thymidine 0
Table 2. Effect of Various Chemicals on Purine-nucleoside phosphorylase
[The enzyme dissolved in 50 mM PIPES buffer, pH 7.0 (10 U/mL) was incubated with each chemical at 30 ℃ for 1 hr.]
-
Chemical Concn.(mM) Residual
activity(%)None - 100 Metal salt 2.0 MgCl2 90.1 CaCl2 96.7 Ba(OAc)2 93.4 FeCl3 73.9 MnCl2 95.0 ZnCl2 77.6 NiCl2 94.1 CuSO4 9.8 Pb(OAc)2 9.1 AgNO3 0.5 HgCl2 0.1 -
Chemical Concn.(mM) Residual
activity(%)N-ethylmaleimide 2.0 91.9 NaF 2.0 90.9 NaN3 20 95.7 EDTA 5.0 96.8 o-Phenanthroline 2.0 98.3 Borate 50 9.0 Iodoacetamide 2.0 98.7 Triton X-100 0.10 % 138.4 Na-cholate 0.10 % 124.4 SDS 0.10 % 0.1 Span 20 0.10 % 128.9
EDTA, Ethylenediaminetetraacetate; SDS, Sodium dodecyl sulfate.
-
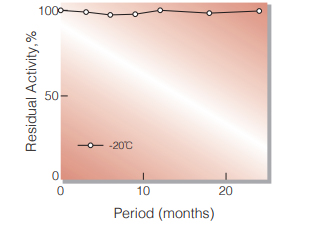
Fig.1. Stability (Powder form)
(kept under dry conditions)
-
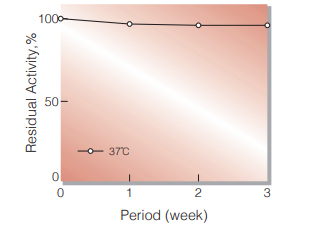
Fig.2. Stability (Powder form)
(kept under dry conditions)
-
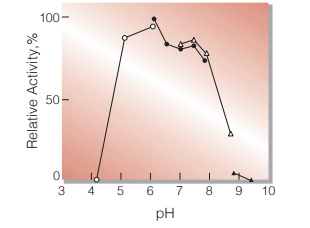
Fig.3. pH-Activity
in 50 mM buffer solution: pH 4-6, acetate; pH 6-8, K-phosphate; pH 7-9, Tris-HCl; pH 9-10, Borate
-
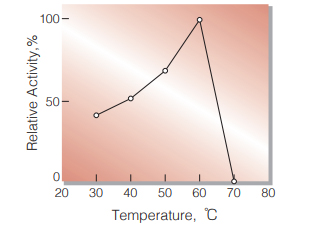
Fig.4. Temperature activity
(in 50 mM K-phosphate buffer, pH 7.7)
-
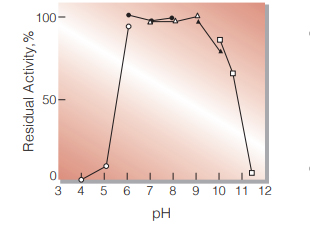
Fig.5. pH-Stability
30 ℃, 16hr-treatment with 50 mM buffer solution: pH 4-6, Acetate; pH 6-8, K-phosphate; pH 7-9, Tris-HCl; pH 9-10, Borate; pH 10-12, Glycine-NaOH. Enzyme concentration: 10 U/mL
-
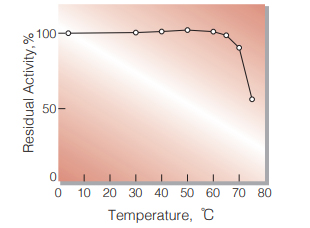
Fig.6. Thermal stability
30 min-treatment with 50 mM K-phosphate buffer, pH 7.7. Enzyme concentration: 1 U/mL
活性測定法(Japanese)
1. 原理

尿酸の生成量を293nmにおける吸光度の変化で測定する。
2.定義
下記条件下で1分間に1マイクロモルの尿酸を生成する酵素量を1単位(U)とする。
3.試薬
- 50mM K-リン酸緩衝液,pH 7.7
- 32mM イノシン水溶液〔85.8mgのイノシン(MW= 268.23)を10mlの蒸留水に加温溶解する〕(4℃ 保存で2週間は使用可能)
- キサンチンオキシダーゼ溶液〔Roche製硫安懸濁 結晶酵素(約20U/ml)を氷冷緩衝液Aで約6U/ml に希釈する〕(用時調製)
酵素溶液:酵素標品を予め氷冷した50mM K-リン酸緩 衝液, pH7.7で溶解し,同緩衝液で0.1~ 1.5U/mlに希釈して氷冷保存する。
4.手順
1.下記反応混液をキュベット(d=1.0cm)に調製し,37℃ で約5分間予備加温する。
| 2.7ml | K-リン酸緩衝液 | (A) |
| 0.20ml | 基質溶液 | (B) |
| 0.10ml | キサンチンオキシダーゼ溶液 | (C) |
2.酵素溶液0.05mlを添加し,ゆるやかに混和後,水を対照に37℃に制御された分光光度計で293nmの吸光度変化を3~4分間記録し,その初期直線部分から1分間当たりの吸光度変化を求める(ΔODtest)。
3.盲検は反応混液①に酵素溶液の代わりに酵素希釈液(50mM K-リン酸緩衝液,pH7.7)を0.05mlを加え, 上記同様に操作を行って,1分間当たりの吸光度変化を求める(ΔODblank)。
5.計算式
-
U/ml =
-
ΔOD/min (ΔOD test−ΔOD blank)×3.05(ml)×希釈倍率
12.5×1.0×0.05(ml)
| = ΔOD/min×4.88×希釈倍率 | |
| U/mg | = U/ml×1/C |
| 12.5 | : 尿酸のミリモル分子吸光係数 (cm2/micromole) |
| 1.0 | : 光路長(cm) |
| C | : 溶解時の酵素濃度(c mg/ml) |
CONTACT
-
For inquiries and cosultations regarding our products, please contact us through this number.
- HEAD OFFICE+81-6-6348-3843
- Inquiry / Opinion
