- HOME
- Enzyme List
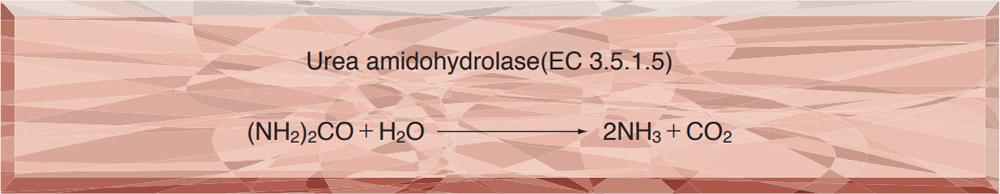
- URH-201 UREASE
URH-201
UREASE from Jack bean
PREPARATION and SPECIFICATION
| Appearance | White amorphous powder, lyophilized | |
|---|---|---|
| Activity | GradeⅡ(-201) 100U/mg-solid or more | |
| Contaminants | Asparaginase | ≤2.0×10-2 % |
| Arginase | ≤2.0×10-3 % | |
| NH4+ | ≤5.0×10-4 μg/U | |
| Stabilizers | EDTA, glutathione, succinate, BSA | |
PROPERTIES
| Stability | Stable at −20 ℃ for at least one year (Fig.1) |
|---|---|
| Molecular weight | approx. 480,000 1) |
| Isoelectric point | 5.0−5.1 1) |
| Michaelis constant | 1.05×10-2 M (Urea) 1) |
| Structure | 8 active sites with SH-groups per the enzyme molecule 2) |
| Inhibitors | Heavy metal ions (Ag+,Hg2+,etc.) |
| Optimum pH | 6.0 (Fig.3) |
| Optimum temperature | 60 ℃ (Fig.4) |
| pH Stability | pH 5.5−8.5 (30 ℃, 17 hr) (Fig.5) |
| Thermal stability | below 50 ℃ (pH 8.0, 60 min) (Fig.6) |
| Effect of various chemicals | (Table.1) |
APPLICATIONS
This enzyme is useful for enzymatic determination of urea in clinical analysis.
ASSAY
Principle

The elimination of NADPH is measured at 340 nm by spectrophotometry.
Unit definition
One unit causes the formation of two micromoles of ammonia per minute under the conditions detailed below.
Method
Reagents
| A. Urea solution | 6.0M (36g of Urea/100 mL of H2O)(Should be prepared fresh) | ||
|---|---|---|---|
| B. Tris-HCl buffer, pH 8.0 | 50mM | ||
| C. α-Ketoglutarate solution | 0.25M (Dissolve 730mg of α-ketoglutarate in 15 mL of H2O, adjust pH to 5.0±0.1 with 5N NaOH and fill up to 20 mL with H2O)(Should be prepared fresh) | ||
| D. NADPH solution | 15mM[Dissolve 136mg of NADPH・Na4・4H2O/10 mL of H2O](Should be prepared fresh) | ||
| E. Working solution (Prepare before use and store on ice) | 69 mL | Tris-HCl buffer | (B) |
| 0.3 mL | α-Ketoglutarate solution | (C) | |
| 1.8 mL | NADPH solution | (D) | |
| 0.9 mL | H2O | ||
| F. GlDH (glutamate dehydrogenase) solution | ca.1,000 U/mL[Toyobo GradeⅡ,GTD-209 (Tris-HCl buffer solution, free from ammonia)] | ||
| G. Enzyme diluent | 10 mM K-phosphate buffer containing 20 mM EDTA and 0.2 % BSA, pH 7.0 | ||
Procedure
1.Prepare the following reaction mixture in a cuvette (d = 1.0 cm) and equilibrate at 37℃ for approximately 5 minutes.
| 2.40 mL | Working solution | (E) |
|---|---|---|
| 0.05 mL | GlDH solution | (F) |
| 0.35 mL | H2O | |
| 0.10 mL | Enzyme solution* |
| Concentration in assay mixture | |
|---|---|
| Tris-HCl buffer | 38 mM |
| Urea | 200 mM |
| α-Ketoglutarate | 0.83 mM |
| NADPH | 0.30 mM |
| EDTA | 0.67 mM |
| GlDH | Approx 17 U/mL |
2.Add 0.10 mL of urea solution (A) and mix by gentle inversion.
3.Record the decrease in optical density at 340 nm against water for 3 to 4 minutes with a spectrophotometer
thermostated at 37 ℃, and calculate the ΔOD per minute from the initial linear portion of the curve (ΔOD
test).
At the same time, measure the blank rate (ΔOD blank) using the same method as in the test except that
the enzyme diluent is added instead of the enzyme solution.
*Dissolve the enzyme preparation in ice-cold enzyme diluent (G) and dilute to 0.07−0.25 U/mL with the same buffer and store on ice.
Calculation
Activity can be calculated by using the following formula :
Volume activity (U/mL) =
-
ΔOD/min (ΔOD test−ΔOD blank)×Vt×df
6.22×2×1.0×Vs
= ΔOD/min×2.41×df
Weight activity (U/mg) = (U/mL)×1/C
| Vt | : Total volume (3.0 mL) |
| Vs | : Sample volume (0.10 mL) |
| 6.22 | : Millimolar extinction coefficient of NADPH at 340 nm (cm2/micromole) |
| 2 | : Factor based on the fact that hydrolysis of one mole of urea is equivalent to oxidation of two moles of NADPH |
| 1.0 | : Light path length (cm) |
| df | : Dilution factor |
| C | : Enzyme concentration in dissolution (c mg/mL) |
REFERENCES
1) J.B.Sumner and D.B.Hand; J.Am.Chem.Soc., 31, 1255 (1929)
2) G.Gorin and C.C.Chin; Biochim.Biophys.Acta., 99, 418 (1965)
3) Rinsho Kagaku Bunseki(Japanese),Ⅱ,p18(M.Kitamura,M.Saito and M.Niwa,ed.)Tokyo Kagaku Dojin,Tokyo (1969)
4) H.G.Schlegel and H.Kaltwasser; Methods of Enzymatic Analysis, Vol.2,p1081 (H.U.Bergmeyer,ed.), Verlag Chemie Weiheim, Acdemic Press, New York-London (1974)
Table 1 Effect of Various Chemicals on UREASE
[The enzyme dissolved in 20mM phosphate buffer, pH 7.0 was incubated with each chemical at 30℃ for 1hr.]
-
Chemical Concn.(mM) Residual
activity(%)None - 100 NaCl 10 96 Na2SO4 10 104 CH3COONa 10 108 Na2HPO4 10 100 Citrate-Na2 10 100 Na2CO3 10 100 Na3BO4 10 104 Na3S2O4 10 108 -
Chemical Concn.(mM) Residual
activity(%)MnCl2 1.0 66 MgCl2 1.0 97 CaCl2 1.0 105 ZnCl2 1.0 104 FeSO4 1.0 94 CuSO4 1.0 99 Ag2SO4 0.1 9 HgCl2 0.1 8
-
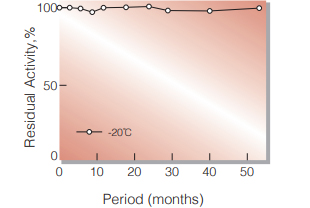
Fig.1. Stability (Powder form)
(kept under dry conditions)
-
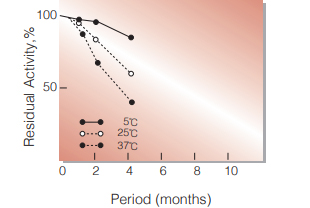
Fig.2. Stability (Powder form)
(kept under dry conditions)
-
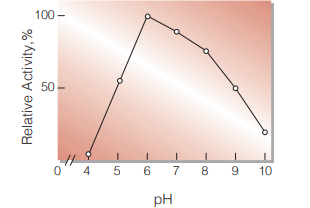
Fig.3. pH-Activity
30℃ in 10mM buffer solution: pH3.0-9.0 Veronal-CH3COONa-HCl; pH9.0-11.0, glycine-NaOH.
-
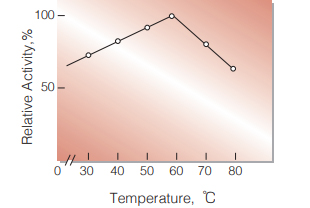
Fig.4. Temperature activity
(in 20mM phosphate buffer,pH7.0)
-
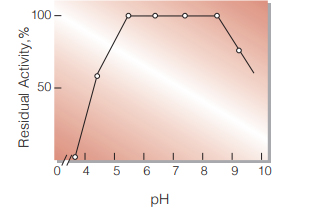
Fig.5. pH-Stability
30℃ ,17hr-treatment with 10mM buffer solution: pH 3.0-9.0, Veronal-CH3COONa-HCI; [pH9.0-11.0: glycine-NaOH
-
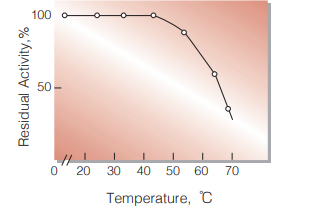
Fig.6. Thermal stability
60min-treatment with 20mM phosphate buffer, pH8.0.
活性測定法(Japanese)
1. 原理

NADPHの減少量を340nmにおける吸光度の変化で測定する。
2.定義
下記条件下で1分間に2マイクロモルのアンモニアを生成する(1マイクロモルの尿素を加水分解する)酵素量を1 単位(U)とする。
3.試薬
- 6.0M 尿素水溶液(36gの尿素を蒸留水に溶解し 100 mLとする)(用時調製)
- 50mM Tris-HCl緩衝液,pH8.0
- 0.25M α-ケトグルタル酸水溶液(730㎎のα-ケトグ ルタル酸を蒸留水約15 mLに溶解し,5N NaOHで pH5.0±0.1に調製後,蒸留水にて20 mLとする)(用時調製)
- 15mM NADPH水溶液〔136㎎のNADPH・Na4・ 4H2Oを蒸留水10 mLに溶解する〕(用時調製)
- 試薬混液(使用直前に調製し,氷冷保存する)
69.0 mL Tris-HCl緩衝液 (B) 0.3 mL α-ケトグルタル酸水溶液 (C) 1.8 mL NADPH水溶液 (D) 0.9 mL 蒸留水 - グルタミン酸脱水素酵素(GlDH)溶液〔東洋紡製, GTD-209(約1,000U/mLに蒸留水で希釈して使用する)〕
酵素溶液:酵素標品を予め氷冷した20mM EDTAと 0.2% BSAを含む10mM K-リン酸緩衝液, pH7.0で溶解し,同緩衝液で0.07~0.25U/mLに希釈して氷冷保存する。
4.手順
1.下記反応混液をキュベット(d=1.0cm)に調製し,37℃ で約5分間予備加温する。
| 2.40 mL | 試薬混液 | (E) |
| 0.05 mL | GlDH溶液 | (F) |
| 0.35 mL | 蒸留水 | |
| 0.10 mL | 酵素溶液 |
2.基質溶液(A)0.10 mLを添加し,ゆるやかに混和後,水を対照に37℃に制御された分光光度計で340nmの吸光度変化を3~4分間記録し,その初期直線部分から1分間当りの吸光度変化を求める(ΔODtest)。
3.盲検は反応混液①に酵素溶液の代わりに酵素希釈液(20mM EDTAを含む10mM K-リン酸緩衝液, pH7.0)を0.10 mL加え,上記同様に操作を行って,1分間当りの吸光度変化を求める(ΔODblank)。
5.計算式
U/mL =
-
ΔOD/min (ΔOD test−ΔOD blank)×3.0(mL)×希釈倍率
6.22×2×1.0×0.10(mL)
| = ΔOD/min×2.41×希釈倍率 | |
| U/mg | = U/mL×1/C |
| 6.22 | : NADPHのミリモル分子吸光係数 (cm2/micromole) |
| 2 | : 酵素反応で1分子の尿素の加水分解に由来するNADPHの酸化は2分子である事による係数 |
| 1.0 | : 光路長(cm) |
| C | : 溶解時の酵素濃度(c mg/mL) |
CONTACT
-
For inquiries and cosultations regarding our products, please contact us through this number.
- HEAD OFFICE+81-6-6348-3843
- Inquiry / Opinion
